Mutation can result in several different types of change in sequences; these can either have no effect, alter the product of a gene, or prevent the gene from functioning properly or completely. One study on genetic variations between different species of Drosophila suggests that if a mutation changes a protein produced by a gene, the result is likely to be harmful, with an estimated 70 percent of amino acid polymorphisms having damaging effects, and the remainder being either neutral or weakly beneficial. Due to the damaging effects that mutations can have on genes, organisms have mechanisms such as DNA repair to prevent mutations.
Mutations can involve large sections of DNA becoming duplicated, usually through genetic recombination. These duplications are a major source of raw material for evolving new genes, with tens to hundreds of genes duplicated in animal genomes every million years. Most genes belong to larger families of genes of shared ancestry. Novel genes are produced by several methods, commonly through the duplication and mutation of an ancestral gene, or by recombining parts of different genes to form new combinations with new functions.
Here, domains act as modules, each with a particular and independent function, that can be mixed together to produce genes encoding new proteins with novel properties. For example, the human eye uses four genes to make structures that sense light: three for color vision and one for night vision; all four arose from a single ancestral gene. Another advantage of duplicating a gene (or even an entire genome) is that this increases redundancy; this allows one gene in the pair to acquire a new function while the other copy performs the original function. Other types of mutation occasionally create new genes from previously noncoding DNA.
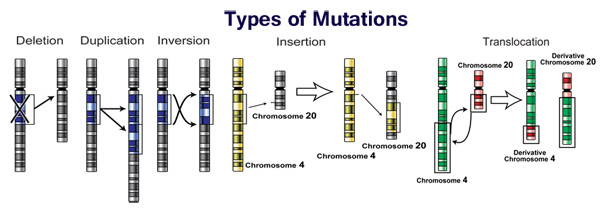
Changes in chromosome number may involve even larger mutations, where segments of the DNA within chromosomes break and then rearrange. For example, in the Homininae, two chromosomes fused to produce human chromosome 2; this fusion did not occur in the lineage of the other apes, and they retain these separate chromosomes. In evolution, the most important role of such chromosomal rearrangements may be to accelerate the divergence of a population into new species by making populations less likely to interbreed, and thereby preserving genetic differences between these populations.
Sequences of DNA that can move about the genome, such as transposons, make up a major fraction of the genetic material of plants and animals, and may have been important in the evolution of genomes. For example, more than a million copies of the Alu sequence are present in the human genome, and these sequences have now been recruited to perform functions such as regulating gene expression. Another effect of these mobile DNA sequences is that when they move within a genome, they can mutate or delete existing genes and thereby produce genetic diversity.
Nonlethal mutations accumulate within the gene pool and increase the amount of genetic variation. The abundance of some genetic changes within the gene pool can be reduced by natural selection, while other “more favorable” mutations may accumulate and result in adaptive changes.
For example, a butterfly may produce offspring with new mutations. The majority of these mutations will have no effect; but one might change the color of one of the butterfly’s offspring, making it harder (or easier) for predators to see. If this color change is advantageous, the chance of this butterfly surviving and producing its own offspring are a little better, and over time the number of butterflies with this mutation may form a larger percentage of the population.
Neutral mutations are defined as mutations whose effects do not influence the fitness of an individual. These can accumulate over time due to genetic drift. It is believed that the overwhelming majority of mutations have no significant effect on an organism’s fitness. Also, DNA repair mechanisms are able to mend most changes before they become permanent mutations, and many organisms have mechanisms for eliminating otherwise permanently mutated somatic cells.
Beneficial mutations can improve reproductive success.
Causes
Spontaneous mutation
Spontaneous mutations on the molecular level can be caused by:
- ·Tautomerism – A base is changed by the repositioning of a hydrogen atom, altering the hydrogen bonding pattern of that base resulting in incorrect base pairing during replication.
- ·Depurination – Loss of a purine base (A or G) to form an apurinic site (AP site).
- ·Deamination – Hydrolysis changes a normal base to an atypical base containing a keto group in place of the original amine group. Examples include C → U and A → HX (hypoxanthine), which can be corrected by DNA repair mechanisms; and 5MeC (5-methylcytosine) → T, which is less likely to be detected as a mutation because thymine is a normal DNA base.
- ·Slipped strand mispairing – Denaturation of the new strand from the template during replication, followed by renaturation in a different spot (“slipping”). This can lead to insertions or deletions.
Induced mutation
Induced mutations on the molecular level can be caused by:
Chemicals
- ·Hydroxylamine NH2OH
- ·Base analogs (e.g. BrdU)
- ·Alkylating agents (e.g. N-ethyl-N-nitrosourea) These agents can mutate both replicating and non-replicating DNA. In contrast, a base analog can only mutate the DNA when the analog is incorporated in replicating the DNA. Each of these classes of chemical mutagens has certain effects that then lead to transitions, transversions, or deletions.
- ·Agents that form DNA adducts (e.g. ochratoxin A metabolites)
- ·DNA intercalating agents (e.g. ethidium bromide)
- ·DNA crosslinkers
- ·Oxidative damage
- ·Nitrous acid converts amine groups on A and C to diazo groups, altering their hydrogen bonding patterns which leads to incorrect base pairing during replication.
Radiation
- ·Ultraviolet radiation (nonionizing radiation). Two nucleotide bases in DNA – cytosine and thymine – are most vulnerable to radiation that can change their properties. UV light can induce adjacent pyrimidine bases in a DNA strand to become covalently joined as a pyrimidine dimer. UV radiation, particularly longer-wave UVA, can also cause oxidative damage to DNA. Mutation rates also vary across species. Evolutionary biologists[citation needed] have theorized that higher mutation rates are beneficial in some situations, because they allow organisms to evolve and therefore adapt more quickly to their environments. For example, repeated exposure of bacteria to antibiotics, and selection of resistant mutants, can result in the selection of bacteria that have a much higher mutation rate than the original population (mutator strains).
Inheritance of Genetic Mutation
A germline mutation gives rise to a constitutional mutation in the offspring, that is, a mutation that is present in every cell. A constitutional mutation can also occur very soon after fertilisation, or continue from a previous constitutional mutation in a parent.
The distinction between germline and somatic mutations is important in animals that have a dedicated germ line to produce reproductive cells. However, it is of little value in understanding the effects of mutations in plants, which lack dedicated germ line. The distinction is also blurred in those animals that reproduce asexually through mechanisms such as budding, because the cells that give rise to the daughter organisms also give rise to that organisms germ line. A new mutation that was not inherited from either parent is called a de novo mutation.
Diploid organisms (e.g. human)contain two copies of each gene – a paternal and a maternal allele. Based on the occurrence of mutation on each chromosome, we may classify mutations into three types.
- · A heterozygous mutation is a mutation of only one allele.
- ·A homozygous mutation is an identical mutation of both the paternal and maternal alleles.
- ·Compound heterozygous mutations or a genetic compound comprises two different mutations in the paternal and maternal alleles.
A wildtype or homozygous non-mutated organism is one in which neither allele is mutated.
Harmful Mutations
If a mutation is present in a germ cell, it can give rise to offspring that carries the mutation in all of its cells. This is the case in hereditary diseases. In particular, if there is a mutation in a DNA repair gene within a germ cell, humans carrying such germ-line mutations may have in increased risk of cancer, such as the list on Wikipedia of inherited human DNA repair gene mutations that increase cancer risk. On the other hand, a mutation may occur in a somatic cell of an organism. Such mutations will be present in all descendants of this cell within the same organism, and certain mutations can cause the cell to become malignant, and thus cause cancer.
A DNA damage can cause an error when the DNA is replicated, and this error of replication can cause a gene mutation that, in turn, could cause a genetic disorder. DNA damages are repaired by the DNA repair system of the cell. Each cell has a number of pathways through which enzymes recognize and repair damages in DNA. Because DNA can be damaged in many ways, the process of DNA repair is an important way in which the body protects itself from disease. Once a DNA damage has given rise to a mutation, the mutation cannot be repaired. DNA repair pathways can only recognize and act on “abnormal” structures in the DNA. Once a mutation occurs in a gene sequence it then has normal DNA structure and cannot be repaired.
Beneficial Mutations
For example, a specific 32 base pair deletion in human CCR5 (CCR5-Δ32) confers HIV resistance to homozygotes and delays AIDS onset in heterozygotes. The CCR5 mutation is more common in those of European descent. One possible explanation of the etiology of the relatively high frequency of CCR5-Δ32 in the European population is that it conferred resistance to the bubonic plague in mid-14th century Europe. People with this mutation were more likely to survive infection; thus its frequency in the population increased. This theory could explain why this mutation is not found in southern Africa, where the bubonic plague never reached. A newer theory suggests that the selective pressure on the CCR5 Delta 32 mutation was caused by smallpox instead of the bubonic plague.
Another example is Sickle cell disease, a blood disorder in which the body produces an abnormal type of the oxygen-carrying substance hemoglobin in the red blood cells. One-third of all indigenous inhabitants of Sub-Saharan Africa carry the gene, because in areas where malaria is common, there is a survival value in carrying only a single sickle-cell gene (sickle cell trait). Those with only one of the two alleles of the sickle-cell disease are more resistant to malaria, since the infestation of the malaria plasmodium is halted by the sickling of the cells which it infests.
Another research from Denmark concludes that blue-eyes are the mutated character of human eyes which were originally brown from around 6,000 to 10,000 years ago. The benign mutation actually effected the OAC2 gene which colorizes our hair and has other functions related to liver e.t.c. So all blue-eyed people share a common ancestor.
Mutation Rate
There are several natural units of time for each of these rates, with rates being characterized either as mutations per base pair per cell division, per gene per generation, or per genome per generation. The mutation rate of an organisms is an evolved characteristic and is strongly influenced by the genetics of each organisms, in addition to strong influence from the environment. The upper and lower limits to which mutation rates can evolve is the subject of ongoing investigation.
Variation in Mutation Rates
If the rate of neutral mutations in a sequence is assumed to be constant (clock-like), and if most differences between species are neutral rather than adaptive, then the number of differences between two different species can be used to estimate how long ago two species diverged (see molecular clock). In fact, the mutation rate of an organism may change in response to environmental stress. For example UV light damages DNA, which may result in error prone attempts by the cell to perform DNA repair.
The human mutation rate is higher in the male germ line (sperm) than the female (egg cells), but estimates of the exact rate have varied by an order of magnitude or more.
In general, the mutation rate in unicellular eukaryotes and bacteria is roughly 0.003 mutations per genome per generation. The highest per base pair per generation mutation rates are found in viruses, which can have either RNA or DNA genomes. DNA viruses have mutation rates between 10−6 to 10−8 mutations per base per generation, and RNA viruses have mutation rates between 10−3 to 10−5 per base per generation. Human mitochondrial DNA has been estimated to have mutation rates of ~3× or ~2.7×10−5 per base per 20 year generation (depending on the method of estimation); these rates are considered to be significantly higher than rates of human genomic mutation at ~2.5×10−8 per base per generation. Using data available from whole genome sequencing, the human genome mutation rate is similarly estimated to be ~1.1×10−8 per site per generation.
The rate for other forms of mutation also differs wildly from point mutations. An individual microsatellite locus often has a mutation rate on the order of 10−4, though this can differ wildly with length.
Mutation in Evolution
Studies have shown that treating RNA viruses such as poliovirus with ribavirin produce results consistent with the idea that the viruses mutated too frequently to maintain the integrity of the information in their genomes. This is termed error catastrophe.
Mutation: Helpful Links
Modern Theories of Evolution: Mutation
Evolution: Library: A Mutation Story
Molecular Evolution, Mutation Size and Gene Pleiotropy: A Geometric Reexamination
Bertram J (2000). “The molecular biology of cancer”. Mol. Aspects Med. 21 (6): 167–223. doi:10.1016/S0098-2997(00)00007-8. PMID 11173079.
Aminetzach YT, Macpherson JM, Petrov DA (2005). “Pesticide resistance via transposition-mediated adaptive gene truncation in Drosophila”. Science 309 (5735): 764–7. doi:10.1126/science.1112699. PMID 16051794.
Burrus V, Waldor M (2004). “Shaping bacterial genomes with integrative and conjugative elements”. Res. Microbiol. 155 (5): 376–86. doi:10.1016/j.resmic.2004.01.012. PMID 15207870.
Sawyer SA, Parsch J, Zhang Z, Hartl DL (2007). “Prevalence of positive selection among nearly neutral amino acid replacements in Drosophila”. Proc. Natl. Acad. Sci. U.S.A. 104 (16): 6504–10. doi:10.1073/pnas.0701572104. PMC 1871816. PMID 17409186.
Hastings, P J; Lupski, JR; Rosenberg, SM; Ira, G (2009). “Mechanisms of change in gene copy number”. Nature Reviews. Genetics 10 (8): 551–564. doi:10.1038/nrg2593. PMC 2864001. PMID 19597530.
Carroll SB, Grenier J, Weatherbee SD (2005). From DNA to Diversity: Molecular Genetics and the Evolution of Animal Design. Second Edition. Oxford: Blackwell Publishing. ISBN 1-4051-1950-0.
Harrison P, Gerstein M (2002). “Studying genomes through the aeons: protein families, pseudogenes and proteome evolution”. J Mol Biol 318 (5): 1155–74. doi:10.1016/S0022-2836(02)00109-2. PMID 12083509.
Orengo CA, Thornton JM (2005). “Protein families and their evolution-a structural perspective”. Annu. Rev. Biochem. 74: 867–900. doi:10.1146/annurev.biochem.74.082803.133029. PMID 15954844.
Long M, Betrán E, Thornton K, Wang W (November 2003). “The origin of new genes: glimpses from the young and old”. Nat. Rev. Genet. 4 (11): 865–75. doi:10.1038/nrg1204. PMID 14634634.
Wang M, Caetano-Anollés G (2009). “The evolutionary mechanics of domain organization in proteomes and the rise of modularity in the protein world”. Structure 17 (1): 66–78. doi:10.1016/j.str.2008.11.008. PMID 19141283.
Bowmaker JK (1998). “Evolution of colour vision in vertebrates”. Eye (London, England) 12 (Pt 3b): 541–7. doi:10.1038/eye.1998.143. PMID 9775215.
Gregory TR, Hebert PD (1999). “The modulation of DNA content: proximate causes and ultimate consequences”. Genome Res. 9 (4): 317–24. doi:10.1101/gr.9.4.317 (inactive 2009-11-14). PMID 10207154.
Hurles M (July 2004). “Gene Duplication: The Genomic Trade in Spare Parts”. PLoS Biol. 2 (7): E206. doi:10.1371/journal.pbio.0020206. PMC 449868. PMID 15252449.
Liu N, Okamura K, Tyler DM (2008). “The evolution and functional diversification of animal microRNA genes”. Cell Res. 18 (10): 985–96. doi:10.1038/cr.2008.278. PMC 2712117. PMID 18711447.
Siepel A (October 2009). “Darwinian alchemy: Human genes from noncoding DNA”. Genome Res. 19 (10): 1693–5. doi:10.1101/gr.098376.109. PMC 2765273. PMID 19797681.
Zhang J, Wang X, Podlaha O (2004). “Testing the Chromosomal Speciation Hypothesis for Humans and Chimpanzees”. Genome Res. 14 (5): 845–51.
Ayala FJ, Coluzzi M (2005). “Chromosome speciation: Humans, Drosophila, and mosquitoes”. Proc. Natl. Acad. Sci. U.S.A. 102 (Suppl 1): 6535–42. doi:10.1073/pnas.0501847102. PMC 1131864. PMID 15851677.
Hurst GD, Werren JH (2001). “The role of selfish genetic elements in eukaryotic evolution”. Nat. Rev. Genet. 2 (8): 597–606. doi:10.1038/35084545. PMID 11483984.
Häsler J, Strub K (2006). “Alu elements as regulators of gene expression”. Nucleic Acids Res. 34 (19): 5491–7. doi:10.1093/nar/gkl706. PMC 1636486. PMID 17020921.
Eyre-Walker A, Keightley PD (August 2007). “The distribution of fitness effects of new mutations”. Nature Reviews Genetics 8 (8): 610–8. doi:10.1038/nrg2146. PMID 17637733.
“Mutation, Mutagens, and DNA Repair”, by Beth A. Montelone, Ph. D., Division of Biology, Kansas State University, 1998
Pfohl-Leszkowicz A, Manderville RA (January 2007). “Ochratoxin A: An overview on toxicity and carcinogenicity in animals and humans”. Mol Nutr Food Res 51 (1): 61–99. doi:10.1002/mnfr.200600137. PMID 17195275.
Kozmin S, Slezak G, Reynaud-Angelin A, Elie C, de Rycke Y, Boiteux S, Sage E (September 2005). “UVA radiation is highly mutagenic in cells that are unable to repair 7,8-dihydro-8-oxoguanine in Saccharomyces cerevisiae”. Proc. Natl. Acad. Sci. U.S.A. 102 (38): 13538–43. doi:10.1073/pnas.0504497102. PMC 1224634. PMID 16157879.
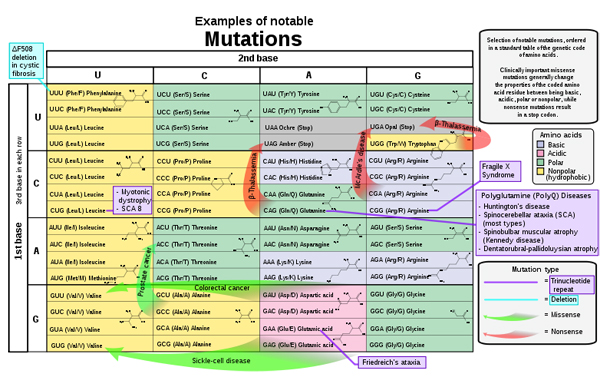
References for the image are found in Wikimedia Commons page at: Commons:File:Notable mutations.svg#References.
Freese, Ernst (April 1959). “THE DIFFERENCE BETWEEN SPONTANEOUS AND BASE-ANALOGUE INDUCED MUTATIONS OF PHAGE T4”. Proc. Natl. Acad. Sci. U.S.A. 45 (4): 622–33. doi:10.1073/pnas.45.4.622. PMC 222607. PMID 16590424.
Freese, Ernst (1959). “The Specific Mutagenic Effect of Base Analogues on Phage T4”. J. Mol. Biol. 1 (2): 87–105. doi:10.1016/S0022-2836(59)80038-3.
Ellis NA, Ciocci S, German J (2001). “Back mutation can produce phenotype reversion in Bloom syndrome somatic cells”. Hum Genet 108 (2): 167–73. doi:10.1007/s004390000447. PMID 11281456.
Charlesworth, D; Charlesworth B, Morgan M T (1995). “The pattern of neutral molecular variation under the background selection model.”. Genetics 141 (4): 1619–32. PMC 1206892. PMID 8601499.
Loewe, L (2006). “Quantifying the genomic decay paradox due to Muller’s ratchet in human mitochondrial DNA.”. Genet Res 87 (2): 133–59. doi:10.1017/S0016672306008123. PMID 16709275.
Peck, J R; Barreau G, Heath S C (1997). “Imperfect genes, Fisherian mutation and the evolution of sex.”. Genetics 145 (4): 1171–99. PMC 1207886. PMID 9093868.
Keightley, P.D.; Lynch M (2003). “Toward a realistic model of mutations affecting fitness.”. Evolution 57 (3): 683–689. JSTOR 3094781?. PMID 12703958.
Barton, N.H.; Keightley P.D. (2002). “Understanding quantitative genetic variation”. Nat Rev Genet 3 (1): 11–21. doi:10.1038/nrg700. PMID 11823787.
